Launching FIREFLY for Full Spectral Fitting
Once your input data is prepared and you have configured FIREFLY with your desired settings, you can start fitting galaxy spectra. Navigate the repository to the Launch section and follow the instructions for your specific dataset.
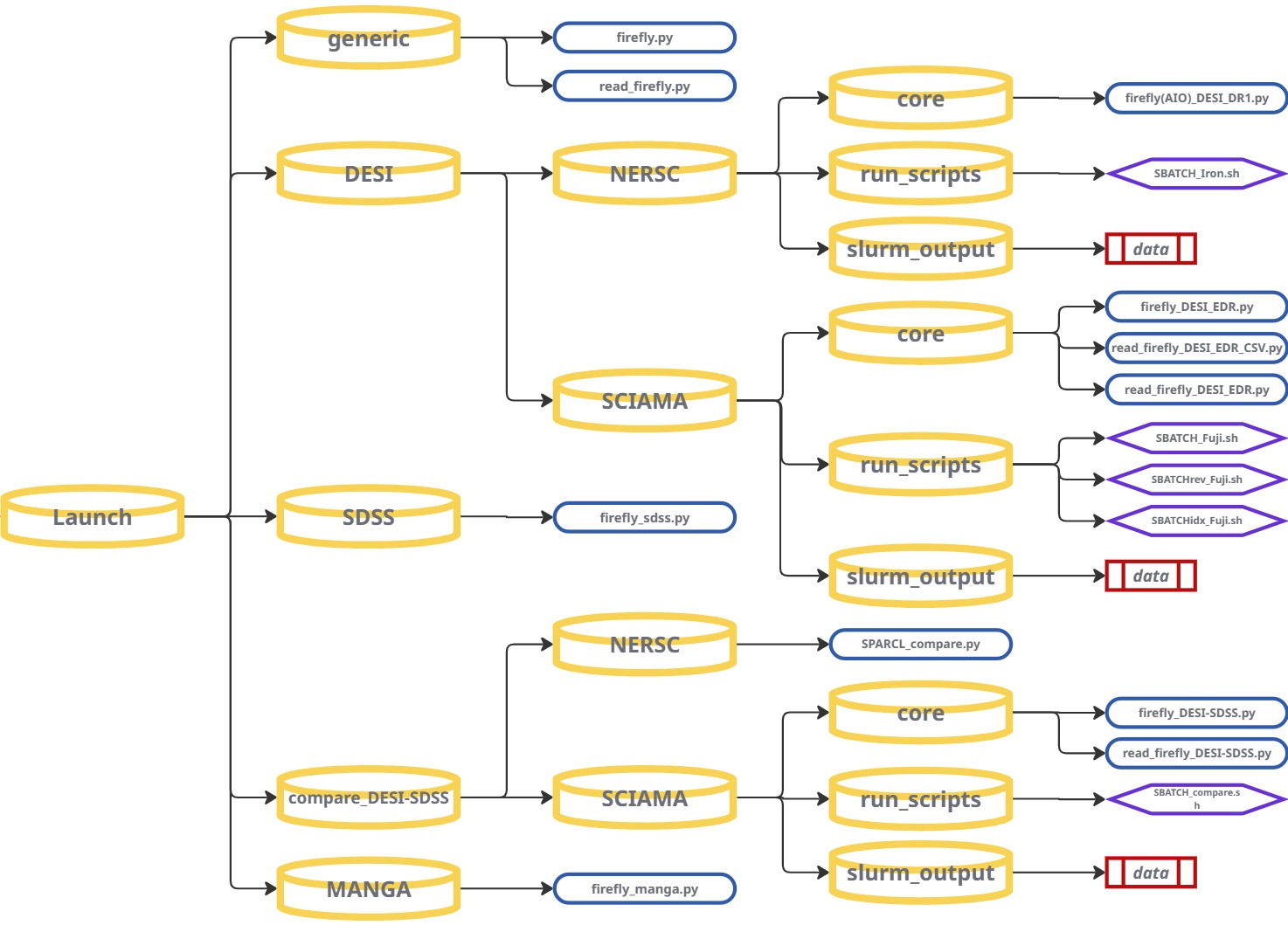
GENERIC — Custom Data / Other Surveys
At its very core, FIREFLY is a Python package that can be run on any
input spectrum, provided the minimal required inputs are given. FIREFLY provides
a base-level initialisation script, firefly.py, that runs the fitting procedure from the most basic setup and initialisation format.
read_firefly.py is also provided to read and plot the output file in its shortest and simplest form.
These generic scripts have been designed to be adapted and redeveloped to read in different data formats you may have, and run FIREFLY on variety of unique data sources. When adapting these base-level FIREFLY scripts to new data, it is recommended to fit only a single galaxy spectrum first. This will help to verify that the input data is being read correctly and the outputs are sensible, before scaling up to larger samples or parallel runs on High Performance Computing (HPC) systems.
Imports — Packages required to launch FIREFLY
FIREFLY is intentionally lightweight and depends mainly on numpy, astropy, and its own internal modules firefly_setup and firefly_models.
# -----------------------------
# firefly.py (core)
# -----------------------------
import sys, os
import time
import numpy as np
# Astropy core I/O + cosmology
from astropy.io import fits
import astropy.cosmology as co
# FIREFLY internals
import firefly_setup as fs
import firefly_models as fm
# -----------------------------
# read_firefly.py (viewer)
# -----------------------------
import sys
import numpy as np
from astropy.io import fits
import matplotlib.pyplot as pltThe key imports that FIREFLY uses are:
-
firefly_setup(imported asfs) - Creates the “spec” container that FIREFLY fits. The module handles; filtering, masking, Milky Way dust correction, instrumental resolutions and all input validation -
firefly_models(imported asfm) - Loads the model library and handles model interpolation, dust grid generation, chi-squared minimisation, PDF sampling for age, metallicity, mass, and writing output tables. -
numpy- FIREFLY spectral data structures are simplenumpy.ndarrayobjects. Used for wavelength grids, flux arrays, loading data and numerical operations. -
astropy.cosmology- FIREFLY embeds thePlanck15cosmology by default and is only required if you want cosmology-anchored stellar population ages. -
astropy.io.fits- Critical for the reading or writing of any input/output*.fitsfile spectral data. Rewuired for the handling of any multi-extension FITS tables (HDUList,BinTableHDU) -
matplotlib.pyplot- Used only inread_firefly.pyfor basic visualisation of spectra + model fits. -
sys,os,time- Standard Python utilities for paths, script timers, and environment-handling.
Additional imports needed when adapting and scaling-up FIREFLY
These are not required by FIREFLY itself, but may become essential when preparing new types of input data or creating parallel runners:
-
argparse- Highly recommended for creating flexible and robust wrapper scripts around FIREFLY -
multiprocessing,concurrent.futures,subprocess- Essential for implementing custom parallelisation strategies and managing multiple processes for distributed computing tasks. -
fcntl- Provides file locking mechanisms, useful for managing concurrent file access in parallel processing. -
pandas- If your results are stored in CSV catalogs or you need fast table joins.
Minimal inputs to FIREFLY
- Wavelength array (Å) (
wavelength) - Flux array (
flux) - default units:flux_units = 1e-17 - Error or inverse-variance array (
error) - Redshift (
redshift) - RA, Dec (
ra,dec) - velocity dispersion (
vdisp) - Instrumental resolution (
r_instrument)
Initialising FIREFLY
Loading a spectrum (example from firefly.py):
input_file = os.path.join(repo_root, "Data", "example_data", "SDSS_ex", "spec-0266-51602-0001.dat")
data = np.loadtxt(input_file, unpack=True)
wavelength = data[0,:]
flux = data[1,:]
error = data[2,:]
restframe_wavelength = wavelength/(1+redshift)
redshift = 0.021275453
ra= 145.89219 ; dec= 0.059372
vdisp = 135.89957
# simple constant resolution example (also in firefly.py)
r_instrument = np.full(len(wavelength), 2000.0)Opening the spectrum inside FIREFLY:
# Define input object to pass data on to firefly modules and initiate run
spec=fs.firefly_setup(input_file,milky_way_reddening=milky_way_reddening, \
N_angstrom_masked=N_angstrom_masked,\
hpf_mode=hpf_mode,data_wave_medium = data_wave_medium)
# Open single spectrum from arrays
spec.openSingleSpectrum(wavelength, flux, error, redshift, ra, dec, vdisp, emlines, r_instrument)Preparing and running the model fit (from firefly.py):
# Prepare model templates
model = fm.StellarPopulationModel(spec, output_file, cosmo, models = model_key, \
model_libs = model_lib, imfs = imfs, \
age_limits = [age_min,age_max], downgrade_models = True, \
data_wave_medium = data_wave_medium, fit_wave_medium=fit_wave_medium,Z_limits = Z_limits, \
suffix=suffix, use_downgraded_models = False, \
dust_law=dust_law, max_ebv=max_ebv, num_dust_vals=num_dust_vals, \
dust_smoothing_length=dust_smoothing_length,max_iterations=max_iterations, \
pdf_sampling=pdf_sampling, flux_units=flux_units)
# Initiate fit
model.fit_models_to_data()Writing the output FITS file:
prihdr = fm.pyfits.Header()
prihdr['FILE'] = os.path.basename(output_file)
prihdu = fm.pyfits.PrimaryHDU(header=prihdr)
tables = [prihdu]
tables.append(model.tbhdu)
complete_hdus = fm.pyfits.HDUList(tables)
complete_hdus.writeto(output_file)Reading + plotting FIREFLY results (read_firefly.py):
# Read in FIREFLY output file (alter as needed using sys.argv or hard-coded paths)
hdul = fits.open(sys.argv[1])
# or
hdul = fits.open("spFly-spec-0266-51602-0001.fits")
data = hdul[1].data
wave = data['wavelength']
flux = data['original_data']
model = data['firefly_model']
# Print out some of the header information
hdul.close()
hdul.info()
print('age: '+str(np.around(10**hdul[1].header['age_lightW'],decimals=2))+' Gyr')
print('[Z/H]: '+str(np.around(hdul[1].header['metallicity_lightW'],decimals=2))+' dex')
print('log M/Msun: '+str(np.around(hdul[1].header['stellar_mass'],decimals=2)))
print('E(B-V): '+str(np.around(hdul[1].header['EBV'],decimals=2))+' mag')
import matplotlib.pyplot as plt
plt.plot(wave, flux)
plt.plot(wave, model)
plt.show()Plotting the star formation history and Z/H distribution
# Plot the star formation history
fig1=plt.figure()
plt.xlim(0,15)
plt.xlabel('lookback time (Gyr)')
plt.ylabel('frequency')
#plt.bar(10**(csp_age),csp_light,width=1,align='center',edgecolor='k',linewidth=2)
plt.bar(10**(csp_age),csp_light,width=1,align='center',alpha=0.5)
plt.scatter(10**(csp_age),csp_light)
plt.show()
# Plot the metallicity distribution
fig2=plt.figure()
plt.xlim(-2,0.5)
plt.xlabel('[Z/H] (dex)')
plt.ylabel('frequency')
#plt.bar(10**(csp_age),csp_light,width=1,align='center',edgecolor='k',linewidth=2)
plt.bar(csp_Z,csp_light,width=0.1,align='center',alpha=0.5)
plt.scatter(csp_Z,csp_light)
plt.show()Implementing command-line arguments with argparse
Instead of editing hardcoded variables for new runs, command-line arguments allow a new FIREFLY pipeline to be completly flexible and scalable. argparse is a widely used Python library and proves very effective when creating quick and easy-to-use wrappers for FIREFLY.
# Import argparse into your new launch wrapper
import argparse
# Define some command-line arguments (example that could be added to firefly.py)
parser = argparse.ArgumentParser(description="Run FIREFLY on a single spectrum")
parser.add_argument("--input", required=True, help="Input spectrum file")
parser.add_argument("--output", required=True, help="Output spFly FITS file")
parser.add_argument("--spec-index", type=int, default=None, help="Index for multi-object")
args = parser.parse_args()
# args.input, args.output, args.spec_index can now be passed trough the pipeline
Example usage:
# Single spectrum
python firefly_WRAPPER.py --input spectrum.fits --output spFly-spectrum.fits
# Multi-object FITS file
python firefly_WRAPPER.py --input desi_chunk.fits --spec-index 42 --output spFly-042.fitsImplementing Parallel Processing - SLURM (HPC cluster execution)
On HPC systems, the SLURM scheduler controls allocations of resources for your submitted jobs and so just like the DESI pipeline custom SBATCH scripts can be created for your new FIREFLY launch wrapper.
Inputs, output, indices and configuration variables can then also be parsed through the pipeline from the SBATCH script.
#!/bin/bash
#SBATCH --job-name=firefly
#SBATCH --cpus-per-task=32
#SBATCH --time=04:00:00
#SBATCH --output=firefly_%j.log
export FF_MAX_WORKERS=$SLURM_CPUS_PER_TASK
# Pass input, output, and optional spec index to the Python script
python firefly_WRAPPER.py \
--input new_survey.fits \
--output spFly_new_results.fits \
--spec-index 42Or process a range of spectra in a loop:
#!/bin/bash
#SBATCH --job-name=firefly_array
#SBATCH --cpus-per-task=32
#SBATCH --array=0-999%10
#SBATCH --time=04:00:00
#SBATCH --output=firefly_%a.log
export FF_MAX_WORKERS=$SLURM_CPUS_PER_TASK
# Use SLURM_ARRAY_TASK_ID to process different spectra in parallel jobs
SPEC_INDEX=$SLURM_ARRAY_TASK_ID
python firefly_WRAPPER.py \
--input new_survey.fits \
--output spFly_${SPEC_INDEX}.fits \
--spec-index $SPEC_INDEXThese are some examples to scale up FIREFLY on new data using parallel processing and parse arguments. But providing FIREFLY is given the required spectral inputs and metadata, custom wrappers can be made for any new galaxy data sources. As seen from the various survey pipelines FIREFLY provides, the code can be tailored and expanded to explore different spectral data on a large-scale.
Tips & Best Practices
- Verify units and calibration: ensure flux is in 10-17 erg/s/cm²/Å and wavelengths are consistently vacuum or air as expected.
- Check redshift-dependent age limits: FIREFLY constrains template ages by the spectrum's redshift, be sure to adjust this if you want to use a different cosmology.
- Monitor job status: use
squeue --mein the HPC terminal to check running/pending SLURM jobs andscancelto abort if needed. - Organise outputs: direct each array job to its own folder or filename prefix (like the DESI pipelines) to avoid any overwriting and clarity between different runs.
- If resources are tight, adjust HPC memory allocations and lower spectra per job or reduce dust smoothing to speed up fitting.
DESI
FIREFLY provides two different pipeline options to fit DESI galaxies. The recommended All-In-One (AIO) launch on NERSC, and the SCIAMA / other HPC method that uses pre-generated FIREFLY-ready FITS files which you then launch FIREFLY upon.
(Note: For the purposes of this workflow we present a DESI-DR1 example for the FIREFLY AIO pipeline and a DESI-EDR example for the SCIAMA / other HPC pipeline. Both wrappers can be easily converted to fit the other data release by changing between the hard-coded data tags: 'iron' for DESI-DR1 and 'fuji' for DESI-EDR.)
Option 1: FIREFLY 'All-in-one' launch scripts on NERSC
If you have a NERSC account you can run the FIREFLY AIO pipeline which retrieves DESI data
directly from the shared filesystem, runs the fitting and produces an output spFly- with the results.
The pipeline is designed to be launched via SLURM using the provided SBATCH script (SBATCH_Iron.sh) which manages job submission and resource allocation.
The --job-name, --account, CPU and memory allocation settings can be adjusted for the number of galaxies you intend to fit as well as managing individual resource allocation.
#!/bin/bash
#!/bin/bash
#SBATCH --job-name=IRON_AIO
#SBATCH --nodes=1
#SBATCH --ntasks=1
#SBATCH --cpus-per-task=128
#SBATCH --mem=256G
#SBATCH --time=2-00:00:00
#SBATCH --constraint=cpu
#SBATCH --qos=regular
#SBATCH --account=desi
#SBATCH --output=../slurm_output/firefly_fit_%j.out
#SBATCH --error=../slurm_output/firefly_fit_%j.err
Define the range of DESI targets to fit. You can modify these values directly in the SBATCH script or pass them as command-line arguments.
# --- Job Configuration ---
# Define the start and end indices for this specific job run.
# This makes it easy to submit different batches.
START_INDEX=2600000
END_INDEX=2700000Make sure to correct the script path with your personal NERSC username before submitting the job to the NERSC SLURM scheduler.
SCRIPT_PATH="/global/cfs/cdirs/desi/users/~~~USERNAME~~~/FIREFLY/Launch/DESI/NERSC/run_scripts/SBATCH_Iron.sh" Submit the job to SLURM using the sbatch command in a NERSC terminal.
cd FIREFLY/Launch/DESI/NERSC/run_scripts
sbatch SBATCH_Iron.sh
SLURM will then return something like:
Submitted batch job 12345678Once the job begins the logs from the nodes will automatically appear in the slurm_output folder as:
firefly_fit_12345678.out
firefly_fit_12345678.err
Usage notes:
- Run a small test first (tens-hundreds of spectra) to confirm the environment and job submission work properly.
- Adjust CPU and memory settings based on the number of spectra to ensure efficient resource usage.
- Monitor progress on the nodes with the job output and error files and examine generated
spFly-*.fitsfiles in your output directory.
Option 2: Launch FIREFLY on SCIAMA / other HPC
If you have already generated FIREFLY-ready FITS files using NERSC or via manual download, you can run FIREFLY on SCIAMA or another HPC system by following these steps:
- Run the
NERSC_fits_create.pyorLocal_fits_create.pyscript to create FIREFLY-ready FITS files. - Transfer the generated FITS files to the target HPC (e.g. SCIAMA) via manually or using
rsync/scp(dependant on HPC permissions) - Run FIREFLY on each block of galaxies SBATCH scripts on the target cluster.
Before submitting a job first select the output mode you want from the initialisation script firefly_DESI_EDR.py
# CSV ONLY setting #
csv_only = False # Set to True to run read_firefly_DESI_EDR_CSV.py instead of read_firefly_DESI_EDR.py- When set to
Truethe pipeline will just produce the master CSV file for the galaxies in the batch block (e.g:DESI_EDR_VAC/master_files/EDR_master_10000-20000.csv). - When set to
Falsethe pipeline will create a new output directory for that galaxy block with the master CSV file, the plotted spectra with best fit models, look back time and metallicty plots for each galaxy. This output directory will have the following stucture (automatically changing file names for the specified galaxy block range):
DESI_EDR_10000-20000/
├── lookback_time_plots_10000-20000/
│ ├── 0_lookback_time_(EDR_10000-20000).png
│ ├── 1_lookback_time_(EDR_10000-20000).png
│ └── ..._lookback_time_(EDR_10000-20000).png
│
├── master_files/
│ └── EDR_master_10000-20000.csv
│
├── metallicity_plots_10000-20000/
│ ├── 0_metallicity_(EDR_10000-20000).png
│ ├── 1_metallicity_(EDR_10000-20000).png
│ └── ..._metallicity_(EDR_10000-20000).png
│
└── spectra_plots_10000-20000/
├── 0_spectrum_(EDR_10000-20000).png
├── 1_spectrum_(EDR_10000-20000).png
└── ..._spectrum_(EDR_10000-20000).png
(Important: Make sure the initialisation script points to the correct input FITS files, the correct Firefly output directory, and that any module loads or environment paths are appropriate for your HPC system).
Now open a terminal on SCIAMA/other HPC system and navigate to the DESI launch scripts for SCIAMA:
cd FIREFLY/Launch/DESI/SCIAMA/run_scripts/
You can list the available batch scripts with:
ls FIREFLY/Launch/DESI/SCIAMA/run_scripts/
SBATCH_Fuji.sh
SBATCHrev_Fuji.sh
SBATCHidx_Fuji.sh
Inside this directory you will find three different batch scripts for launching FIREFLY on DESI-EDR. These exist as options to help avoid issues such as out-of-memory (OOM) kills, variation in node memory availability, and different execution behaviours between HPC systems.
- SBATCH_Fuji.sh: Standard forward-order processing
- Splits the DESI galaxy block into CPU-sized chunks.
- Fits each galaxy from index 0 → last index within each chunk.
- Recommended for normal, stable FIREFLY runs on SCIAMA.
- Best used when memory pressure is predictable and low.
- SBATCHrev_Fuji.sh: Reverse-order processing
- Processes galaxies in a reverse index order (last → first).
- Helps avoid OOM kills caused by bad memory clustering or model ordering.
- Useful when the forward-order processing misses some galaxies on certain nodes.
- SBATCHidx_Fuji.sh: Missing-galaxy recovery
- Scans the out master CSV file for any missing or failed galaxy fits.
- Creates a list of only the missing indices and reruns FIREFLY for them.
- Ideal for cleanup after large runs or for OOM-kill recovery.
- Ensures the final dataset is complete to the best of FIREFLY's ability, meaning the missing fits left are usually as a result of instrument related errors.
Be sure to adapt the job configuration settings to match your HPC system's resources and job requirements:
#!/bin/bash
#SBATCH --job-name=desifirefly
#SBATCH --partition=sciama3-5.q
#SBATCH --mail-type=ALL
#SBATCH --mail-user=up2013158@myport.ac.uk
#SBATCH --ntasks=32
#SBATCH --cpus-per-task=1
#SBATCH --mem-per-cpu=10G
To adjust the size of the galaxy block for different numbers of galaxies, modify the spectra_end (number of galaxies in fits file) and chunk_size (number of galaxies divided by number of task or CPUs) variables in the script:
spectra_start=0
spectra_end=10000
# Export the file range for output directory naming
export DESI_FILE_RANGE=$RANGE
chunk_size=313
Make sure to correct the paths in the script with your personal SCIAMA/other HPC username before submitting the job to the SLURM scheduler:
export PYTHONPATH='/users/~~~USERNAME~~~/FIREFLY/python:$PYTHONPATH'
export DESI_INPUT_FILE=$INPUT_FILE
cd /users/~~~USERNAME~~~/FIREFLY
Finally launch FIREFLY on SCIAMA or another HPC system, submitting the job to SLURM using the sbatch command the HPC terminal:
sbatch SBATCH_Fuji.sh DESI_EDR_10000-20000.fits
.png)
SDSS
FIREFLY for SDSS is driven from a single initialisation script, firefly_sdss.py. The script reads one SDSS FITS input, runs the fitting procedure and writes a single FIREFLY output file.
Run command
From the launch directory containing firefly_sdss.py run:
python firefly_sdss.pyInput (what the script expects)
The script opens the FITS and extracts the arrays/headers shown here (these must be present in the input file):
input_file='example_data/spec-0266-51602-0001.fits'
hdul = fits.open(input_file)
# extracted by the script
redshift = hdul[2].data['Z'][0]
wavelength = 10**hdul[1].data['loglam']
flux = hdul[1].data['flux']
error = hdul[1].data['ivar']**(-0.5)
ra = hdul[0].header['RA']
dec = hdul[0].header['DEC']
vdisp = hdul[2].data['VDISP'][0]Output (where results are written)
The script creates an output folder (if missing) and writes a single FITS file named from the input basename:
outputFolder = os.path.join( os.getcwd(), 'output')
output_file = os.path.join( outputFolder , 'spFly-' + os.path.basename( input_file )[0:-5] ) + ".fits"If the output file already exists the script prompts before overwriting:
if os.path.isfile(output_file):
answer = input('** Do you want to continue? (Y/N)')
if (answer=='N' or answer=='n'):
sys.exit()
os.remove(output_file)Reading FIREFLY outputs
The FIREFLY result FITS produced by the SDSS script can be inspected with the provided read_firefly.py utility. Run it on the output file:
python read_firefly.py output/spFly-.fits The read_firefly.py script (usage shown below) opens the FITS, prints the primary/header info and derived quantities (light-weighted age, [Z/H], stellar mass, E(B-V)) and plots the original spectrum and best-fit model. It also plots the SSP contributions as lookback-time and metallicity histograms.
# read_firefly.py — behaviour (summary of what it does)
hdul = fits.open(sys.argv[1])
data = hdul[1].data
wave = data['wavelength']
flux = data['original_data']
model = data['firefly_model']
# prints header, age, metallicity, stellar mass, E(B-V)
# plots spectrum + model and two bar plots: lookback-time (age) and [Z/H] histogramsScaling / batch notes
The SDSS initialization script processes the single input file specified by input_file. There is no built-in loop or multiprocessing in this file — to process many SDSS spectra you must invoke the script once per input file (the script will produce one spFly-*.fits result per invocation). Each resulting FITS can be inspected with read_firefly.py.
Run flow & final message
When the run finishes the script writes the FITS and prints a completion time, for example:
complete_hdus.writeto(output_file)
print()
print ("Done... total time:", int(time.time()-t0) ,"seconds.")
print()DESI ↔ SDSS — Dual launch & comparison pipeline
To compare the same galaxies from DESI and SDSS this pipelines uses the SPARCL (SPectra Analysis and Retrievable Catalog Lab) client to match galaxies from the two surveys then runs FIREFLY on both data sets at once to produce side-by-side comparison fits of matched galaxies from DESI and SDSS. The pipeline queries the SPARCL Python client to quickly retrive the spectra from each survey, please refer to Juneau et al. 2024 when using the software and the example notebook by Ragadeepika Pucha (U.Utah), Stéphanie Juneau (NOIRLab), and the Astro Data Lab Team.
Step 1 - match DESI + SDSS galaxies on NERSC (SPARCL)
Run the retrieval script sparcl_compare.py on NERSC. The script:
- Queries SPARCL for SDSS and DESI spectra within a specified RA/Dec/redshift window set near the top of the script (e.g.
RA_MIN/RA_MAX,DEC_MIN/DEC_MAX,Z_MIN/Z_MAX). - Matches objects on-sky using
search_around_skywith a maximum tolerance set byMATCH_RADIUS_ARCSEC(default0.5arcsec) to produce SDSS ↔ DESI pairs. - Retrieves the matched spectra and resamples if necessary (resampling to a linear base grid when wave arrays differ so firefly can fit comparitively), then trims leading/trailing zero/NaN/ivar==0 pixels for each spectrum and drops spectra with fewer than
MIN_GOOD_PIXELS. - Writes two FIREFLY-ready FITS files:
<~~~SAMPLE~~~>_sdss_dr16.fitsand<~~~SAMPLE~~~>_desi_dr1.fits.
Key settings (from the sparcl_compare.py script):
# matching and limits
MATCH_RADIUS_ARCSEC = 0.5
LIMIT = 5000
OUT_PREFIX = "greatwall_sample"
# resampling / trimming
FORCE_RESAMPLE = False
MIN_GOOD_PIXELS = 10
# optional fastspec vdisp mapping:
FASTSPEC_PATH = "/global/cfs/cdirs/desi/public/edr/vac/edr/fastspecfit/fuji/v3.2/catalogs/fastspec-fuji.fits"Ensure the a python environment with sparclclient is installed and run the SPARCL retrieval script on NERSC:
python sparcl_compare.pyAs we use the SPARCL client the script is quick and will output the following FIREFLY-ready FITS files:
greatwall_sample_sdss_dr16.fits
greatwall_sample_desi_dr1.fitsStep 2 - transfer the sample FITS files to the target HPC (e.g. SCIAMA)
Place the two FITS into the sample data folder expected by the compare SBATCH wrapper on the target cluster. The example SBATCH file expects the files to be in:
FIREFLY/firefly/Data/DESI-SDSS_samples/Note: the transfer step could instead be done on NERSC (i.e. run the next stage on NERSC, another HPC or locally) - the compare pipeline just needs read-access to the two FITS files from the sample.
Step 3 - launch FIREFLY DESI ↔ SDSS comparative fitting on the HPC (e.g. SCIAMA)
The 'compare' workflow on the HPC is implemented by:
firefly_DESI-SDSS_compare.py- loads the two matched FITS (DESI + SDSS), loops over a the specifed index range and runs FIREFLY on each matched pair, producing temporary per-spectrum output FITS files for DESI and SDSS, then calls the read script to create comparison plots and update master CSVs.read_firefly_DESI-SDSS_compare.py- analyses the two FIREFLY per-spectrum outputs, extracts the derivered steller population parameters, appends/updates the master CSVs, and produces the combined comparison figures. It stores outputs under a sample-specific base directory.SBATCH_compare.sh- orchestrates splitting the galaxy indices into chunks and launching parallelsruntasks forfirefly_DESI-SDSS_compare.py, submitting the jobs to SLURM.
The SBATCH_compare.sh script sets the required environment variables DESI_INPUT_FILE and SDSS_INPUT_FILE from the sample name you provide. It expects the two matched files to be named as ${SAMPLE}_sample_desi_dr1.fits and ${SAMPLE}_sample_sdss_dr16.fits inside DESI-SDSS_sample_data.
Finally launch the FIREFLY comparative fitting pipeline on SCIAMA/other HPC, submitting the job to SLURM using the sbatch command the HPC terminal:
# run without wavelength cut
sbatch SBATCH_compare.sh greatwall
# run with wavelength cut mode (SDSS-driven crop -> crop DESI to same range)
sbatch SBATCH_compare.sh greatwall cutThere is two method choices for the comparative fitting, specifed with the optional cut varible in the terminal SBATCH job submission. The cut selector defines the treatment of wavelength cropping (CUT_MODE):
# in firefly_DESI-SDSS_compare.py
cut_mode = os.environ.get('CUT_MODE', '').lower()
if cut_mode == 'cut':
# compute min/max valid SDSS wavelength (non-zero)
# crop SDSS to that range and crop DESI to the same min/max (no resampling)
# (prints: "Applied wavelength cut: min - max Å")
else:
# no cut applied; use full wavelength ranges
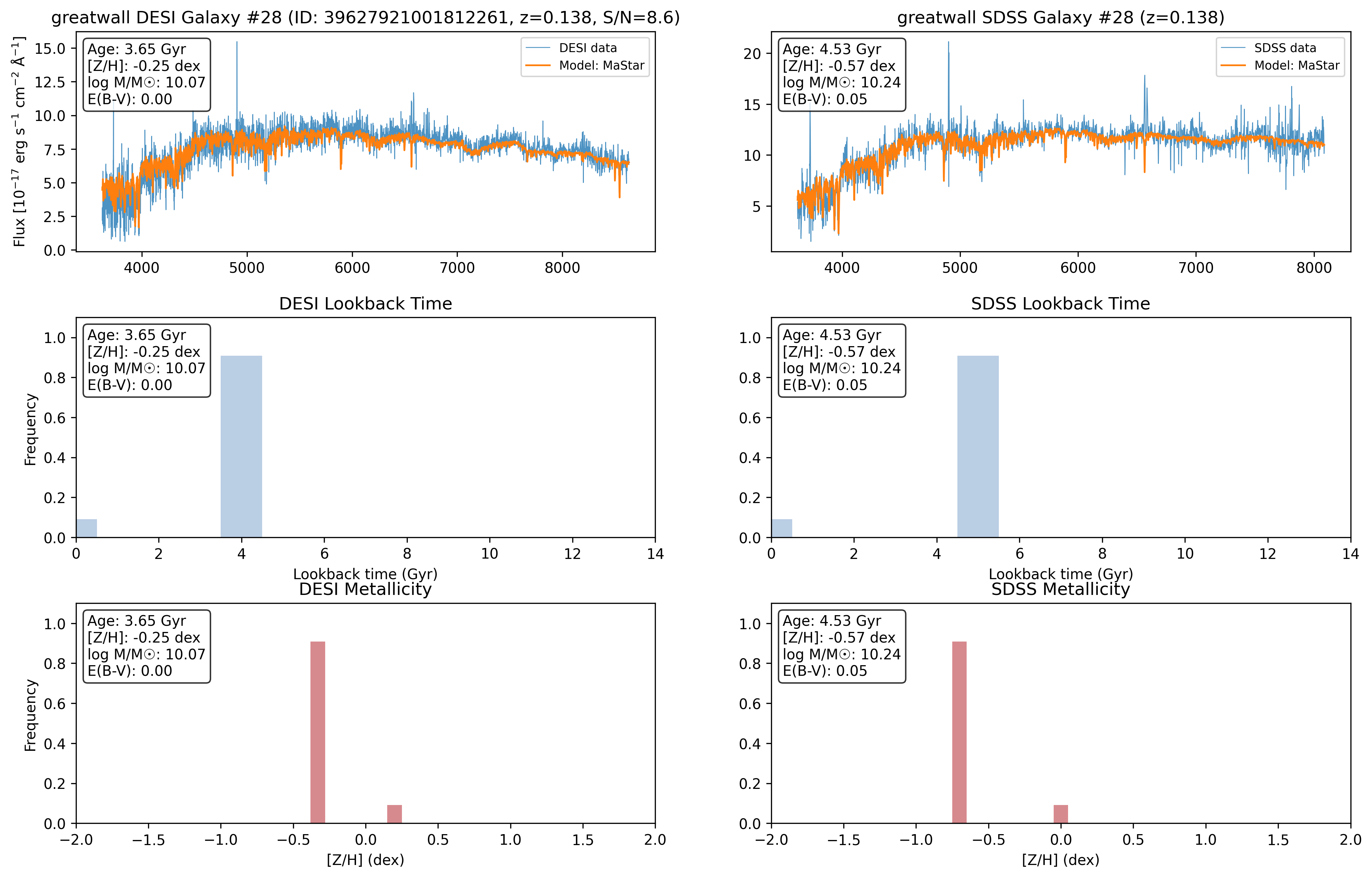
After the run completes, inspect the output directory (greatwall_compare_NoCut) and use the analyse_compare_results.py script in the analysis folder to generate summary plots and statistics comparing the two surveys.
(Note: If you need to install SPARCL client onto the python enviroment you use on NERSC install via pip install sparclclient and consult the SPARCL client documentation for more infomation)
MaNGA
FIREFLY uses a single MaNGA initialization script which can fit a single Voronoi bin or (optionally) all bins via multiprocessing. Set the plate/IFU and either a single bin_number or 'all'.
Run command
From the launch MaNGA directory containing firefly_manga.py run:
python firefly_manga.pyInput / basic configuration
Paths and identifiers are set inside the script:
# Paths to input files
manga_data_dir = os.path.join(repo_root, "Data", "example_data", "MANGA_ex")
logcube_dir = manga_data_dir
maps_dir = manga_data_dir
dap_file = os.path.join(manga_data_dir, "dapall-v3_1_1-3.1.0.fits")
suffix = ""
# Define plate number, IFU number, bin number
plate = 8080
ifu = 12701
bin_number = 0 #'all' if entire IFU, otherwise bin number
Output layout
The script creates a per-plate/IFU output directory and writes one FITS per bin using a filename pattern:
output_dir = os.path.join(repo_root, "Results", "manga")
direc = os.path.join(output_dir,str(plate),str(ifu))
# per-bin output inside f()
output_file = direc + '/spFly-' + str(plate) + '-' + str(ifu) + '-bin' + str(i)Running all bins (multiprocessing)
The script reads the MAPS file to discover unique bin IDs and — if bin_number == 'all' — launches a multiprocessing pool across detected CPU cores:
maps_header = fits.open(maps)
unique_bin_number = list(np.unique(maps_header['BINID'].data)[1:])
print('Number of bins = {}'.format(len(unique_bin_number)))
if bin_number=='all':
N = f_mp(unique_bin_number)
else:
f(bin_number)The multiprocessing wrapper maps the per-bin function f across all bins using multiprocessing.cpu_count():
def f_mp(unique_bin_number):
if __name__ == '__main__':
pool = Pool(processes=multiprocessing.cpu_count())
pool.map(f, unique_bin_number)
pool.close()
pool.join()The read_firefly.py script opens the given FITS, prints headers, prints derived values (light-weighted age, [Z/H], stellar mass, E(B-V)), plots spectrum vs model and shows SSP lookback-time and metallicity bar plots for the fitted result.
Scaling notes
- To process the whole IFU set
bin_number = 'all'- the script will then use multiprocessing to runfacross the bins discovered in the MAPS file. - Outputs are organised under
output/manga/<plate>/<ifu>/with onespFly-...FITS per bin; each output can be inspected withread_firefly.py.
Run flow & final message
Each bin is prepared via fs.firefly_setup, model templates are prepared and model.fit_models_to_data() is called. The script prints the total wall time at the end:
# per-bin fit (summary)
spec = fs.firefly_setup(maps, milky_way_reddening=milky_way_reddening, \
N_angstrom_masked=N_angstrom_masked, hpf_mode=hpf_mode, \
data_wave_medium=data_wave_medium)
spec.openMANGASpectrum(logcube, dap_file, galaxy_bin_number, plate, ifu, emlines, mpl=mpl)
model = fm.StellarPopulationModel(spec, output_file, cosmo, models = model_key, ...)
model.fit_models_to_data()
# final print
print('Time to complete: ', (time.time())-t0)